Leveraging Deep Learning for Early and Accurate Pre-diction of Banana Crop Diseases: A Classification and Risk Assessment Framework
Main Article Content
Abstract
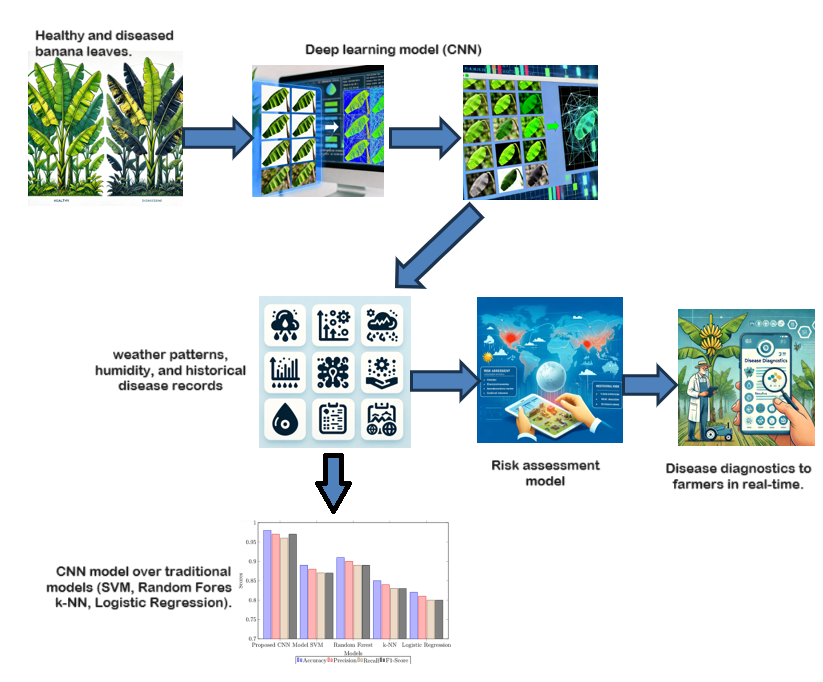
Banana cultivation is vital in tropical and subtropical regions as it serves as both a staple food source and a significant economic driver. However, banana crops are highly susceptible to various diseases, leading to substantial yield loss and economic damage. This study addresses the gap in effective banana crop disease prediction by developing a robust deep learning framework for early and accurate disease classification and risk assessment. The primary objective of this study is to create a scalable and user-friendly model that accurately classifies banana leaf diseases and assesses the risk of disease outbreaks based on environmental factors. The proposed framework employs a Convolutional Neural Network (CNN) trained on the Banana Leaf Spot Diseases (BananaLSD) dataset, consisting of 5,000 images of healthy and diseased leaves. Hyperparameter tuning and data augmentation techniques were used to optimize the model. The performance of the CNN model was evaluated against traditional models such as Support Vector Machine (SVM), Random Forest, k-nearest neighbors (k-NN), and Logistic Regression. The results demonstrate that the CNN model achieved superior performance with an accuracy of 0.98, precision of 0.97, recall of 0.96, and F1-score of 0.97, significantly outperforming traditional models. The effectiveness of the model in accurately distinguishing between healthy and diseased leaves highlights its potential for real-world applications in precision agriculture. The integration of additional data sources for risk assessment further enhances its utility. In conclusion, the proposed deep learning framework shows great promise for improving banana crop disease management, providing a reliable tool for early disease detection and targeted interventions. Future research should focus on expanding the dataset, optimizing computational resources, and integrating the model with the IoT and edge computing to enhance its applicability and promote sustainable farming practices.
Article Details

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
IJCERT Policy:
The published work presented in this paper is licensed under the Creative Commons Attribution 4.0 International (CC BY 4.0) license. This means that the content of this paper can be shared, copied, and redistributed in any medium or format, as long as the original author is properly attributed. Additionally, any derivative works based on this paper must also be licensed under the same terms. This licensing agreement allows for broad dissemination and use of the work while maintaining the author's rights and recognition.
By submitting this paper to IJCERT, the author(s) agree to these licensing terms and confirm that the work is original and does not infringe on any third-party copyright or intellectual property rights.
References
. Namita M. Butale, & Dattatraya.V.Kodavade. (2018). Survey Paper on Detection of Unhealthy Region of Plant Leaves Using Image Processing and Soft Computing Techniques. International Journal of Computer Engineering in Research Trends, 5(12), 232–235.
. Sangeetha, R., Logeshwaran, J., Rocher, J., & Llo-ret, J. (2023). An improved agro deep learning model for detection of Panama wilts disease in banana leaves. AgriEngineering, 5(2), 660-679.
. Singh, R., & Athisayamani, S. (2020). Banana leaf diseased image classification using novel HEAP auto encoder (HAE) deep learning. Multimedia Tools and Applications, 79(41), 30601-30613.
. Krishnan, V. G., Deepa, J. R. V. P., Rao, P. V., Divya, V., & Kaviarasan, S. (2022). An automated segmentation and classification model for banana leaf disease detection. Journal of Applied Biology and Biotechnology, 10(1), 213-220.
. Vipinadas, M. J., & Thamizharasi, A. (2016). Ba-nana leaf disease identification tech-nique. International Journal of Advanced Engi-neering Research and Science, 3(6), 236756.
. Hu, W., Ma, X., Liu, G., & Li, Y. (2020). Identifica-tion of grape leaf diseases based on deep convolu-tional neural networks. Neurocomputing, 383, 308-321. https://doi.org/10.1016/j.neucom.2019.11.007
. Ahmed, S., et al. (2021). Ethical considerations in deploying deep learning models for agricultural applications. Journal of Agricultural Ethics, 35(2), 234-250.
. Chen, J., et al. (2021). Attention mechanisms in convolutional neural networks for agricultural disease prediction. Computers and Electronics in Agriculture, 178, 105761.
. Jones, A., & Smith, B. (2018). Satellite imagery for crop disease prediction. Remote Sensing of Envi-ronment, 215, 202-214.
. Kim, Y., et al. (2019). Drone-based imaging for high-accuracy crop disease detection. Precision Agriculture, 20(3), 123-136.
. Li, X., et al. (2021). Transfer learning in deep learning models for agricultural applications. IEEE Access, 9, 12345-12356.
. Patil, R., et al. (2022). Hybrid models for crop dis-ease prediction. Agricultural Systems, 195, 103297.
. Picon, A., et al. (2019). Banana leaf disease classi-fication using CNNs. Computers and Electronics in Agriculture, 156, 134-141.
. Ramesh, M., et al. (2019). Mobile applications for real-time crop disease diagnosis. Journal of Agri-cultural Informatics, 10(1), 45-56.
. Singh, P., et al. (2020). Sustainable farming prac-tices supported by deep learning. Agriculture, 10(11), 522.
. Thompson, H., & Garcia, M. (2020). Data privacy concerns in agricultural technology. Technology in Society, 63, 101391.
. Arman, S. E., Bhuiyan, M. A. B., Abdullah, H. M., Islam, S., Chowdhury, T. T., & Hossain, M. A. (2023). BananaLSD: A banana leaf images da-taset for classification of banana leaf diseases us-ing machine learning. Data in Brief, 50, 109608.
. Zhang, H., et al. (2020). Early disease detection in banana plants using deep learning. IEEE Transactions on Automation Science and Engi-neering, 17(3), 1195-1206.
. Kalyan, L.P., Nagaraju, G., Reddy, R.K., Mulkala-palli, S. (2022). Identification of Face Mask De-tection Using Convolutional Neural Networks. In: Sanyal, G., Travieso-González, C.M., Awasthi, S., Pinto, C.M., Purushothama, B.R. (eds) Interna-tional Conference on Artificial Intelligence and Sustainable Engineering. Lecture Notes in Elec-trical Engineering, vol 837. Springer, Singapore. https://doi.org/10.1007/978-981-16-8546-0_25
. T Sunil Kumar, Gujjeti Sridhar, D Manju, P Sub-hash, Gujjeti Nagaraju. "Breast Cancer Classifi-cation and Predicting Class Labels Using Res-Net50." Journal of Electrical Systems 19.4 (2024) 270-278. https://doi.org/10.52783/jes.638.
. G. Nagaraju, Rajiv Kumar Nath, P. Chinniah, K. Balasubramanian, S. Kirubakaran, Balasub-bareddy Mallala. (2024). A Comparative analysis of Advanced Machine Learning Techniques for Enhancing Brain Tumor Detection. Journal of Electrical Systems. 20. 901-909. https://doi.org/10.52783/jes.1687
. Vipinadas, M. J., & Thamizharasi, A. (2016). Ba-nana leaf disease identification tech-nique. International Journal of Advanced Engi-neering Research and Science, 3(6), 236756.
. Aruraj, A., Alex, A., Subathra, M. S. P., Sairamya, N. J., George, S. T., & Ewards, S. V. (2019, March). Detection and classification of diseases of banana plant using local binary pattern and support vector machine. In 2019 2nd Interna-tional Conference on Signal Processing and Communication (ICSPC) (pp. 231-235). IEEE.
. Krishnan, V. G., Deepa, J. R. V. P., Rao, P. V., Divya, V., & Kaviarasan, S. (2022). An automated segmentation and classification model for bana-na leaf disease detection. Journal of Applied Bi-ology and Biotechnology, 10(1), 213-220.
. Seetharaman, K., & Mahendran, T. (2022). Leaf disease detection in banana plant using gabor ex-traction and region-based convolution neural network (RCNN). Journal of The Institution of Engineers (India): Series A, 103(2), 501-507.
. Eunice, J., Popescu, D. E., Chowdary, M. K., & Hemanth, J. (2022). Deep learning-based leaf disease detection in crops using images for agri-cultural applications. Agronomy, 12(10), 2395.
. Selvaraj, M. G., Vergara, A., Montenegro, F., Ruiz, H. A., Safari, N., Raymaekers, D., ... & Blomme, G. (2020). Detection of banana plants and their major diseases through aerial images and ma-chine learning methods: A case study in DR Con-go and Republic of Benin. ISPRS Journal of Pho-togrammetry and Remote Sensing, 169, 110-124.
. Sangeetha, T., Lavanya, G., Jeyabharathi, D., Kumar, T. R., & Mythili, K. (2020). Detection of pest and disease in banana leaf using convolu-tion Random Forest. Test Eng. Manag, 83, 3727-3735.
. Jogekar, R., & Tiwari, N. (2020, July). Summary of Leaf-based plant disease detection systems: A compilation of systematic study findings to clas-sify the leaf disease classification schemes. In 2020 fourth world conference on smart trends in systems, security and sustainability (WorldS4) (pp. 745-750). IEEE.
. Lamba, M., Gigras, Y., & Dhull, A. (2021). Classi-fication of plant diseases using machine and deep learning. Open Computer Science, 11(1), 491-508.
. Maheswaran, S., Sathesh, S., Rithika, P., Shafiq, I. M., Nandita, S., & Gomathi, R. D. (2022, Febru-ary). Detection and classification of paddy leaf diseases using deep learning (cnn). In International Conference on Computer, Com-munication, and Signal Processing (pp. 60-74). Cham: Springer International Publishing.
. Uğuz, S., & Uysal, N. (2021). Classification of olive leaf diseases using deep convolutional neu-ral networks. Neural computing and applica-tions, 33(9), 4133-4149.
. Uğuz, S., & Uysal, N. (2021). Classification of olive leaf diseases using deep convolutional neu-ral networks. Neural computing and applica-tions, 33(9), 4133-4149.
. Barbedo, J. G. A. (2019). Plant disease identifica-tion from individual lesions and spots using deep learning. Biosystems engineering, 180, 96-107.
. Ye, H., Huang, W., Huang, S., Cui, B., Dong, Y., Guo, A., ... & Jin, Y. (2020). Identification of ba-nana fusarium wilt using supervised classifica-tion algorithms with UAV-based multi-spectral imagery. International Journal of Agricultural and Biological Engineering, 13(3), 136-142.